The CompBio Lab
People
- Anna Ritz (PI)
- I joined the Reed College Biology Department in the Fall of 2015. For more information, see the home page, my CV, or the links on the right-side banner.
- Current Members
- Pramesh Singh, postdoctoral researcher
- Alumni
- Nick Franzese, post-bac researcher
- Alexander King, post-bac researcher
- Amy Rose Lazarte, post-bac researcher
- Darsh Mandera, high schooler
- Ramin Neshati, visiting scholar
- Gabe Preising, post-bac researcher
- Alex Richter, post-bac researcher
- Tobias Rubel, post-bac researcher
- Ibrahim Youssef, postdoctoral researcher
- Summer 2022: Hannah Kuder (Math/Physics), Henry Jacques (Biochemistry), Li Yi (Math/CS), Nina Young (CS/Art), Larry Zeng (Math/CS)
- Summer 2020: Jiarong Li (Math/CS). Alex Richter (Math/CS), Aryeh Stahl (Math/CS), Larry Zeng (Math/CS), Frank Zhang (Biology)
- Summer 2019: Tayla Isensee (Biology), Tunc Kose (Math/CS), Jiarong Li (Math/CS), Karl Young (Biology)
- Summer 2018: Miriam Bern (Biology), Alexander King (Neuroscience), Sol Taylor Brill (Biology), Kathy Thompson (CS), Usman Hafeez (Math)
- Summer 2017: Nick Egan (Sociology), Giorlando Ramirez (Economics), Yurel Watson (CS)
- Summer 2016: Karl Menzel (Biology)
Group Photos
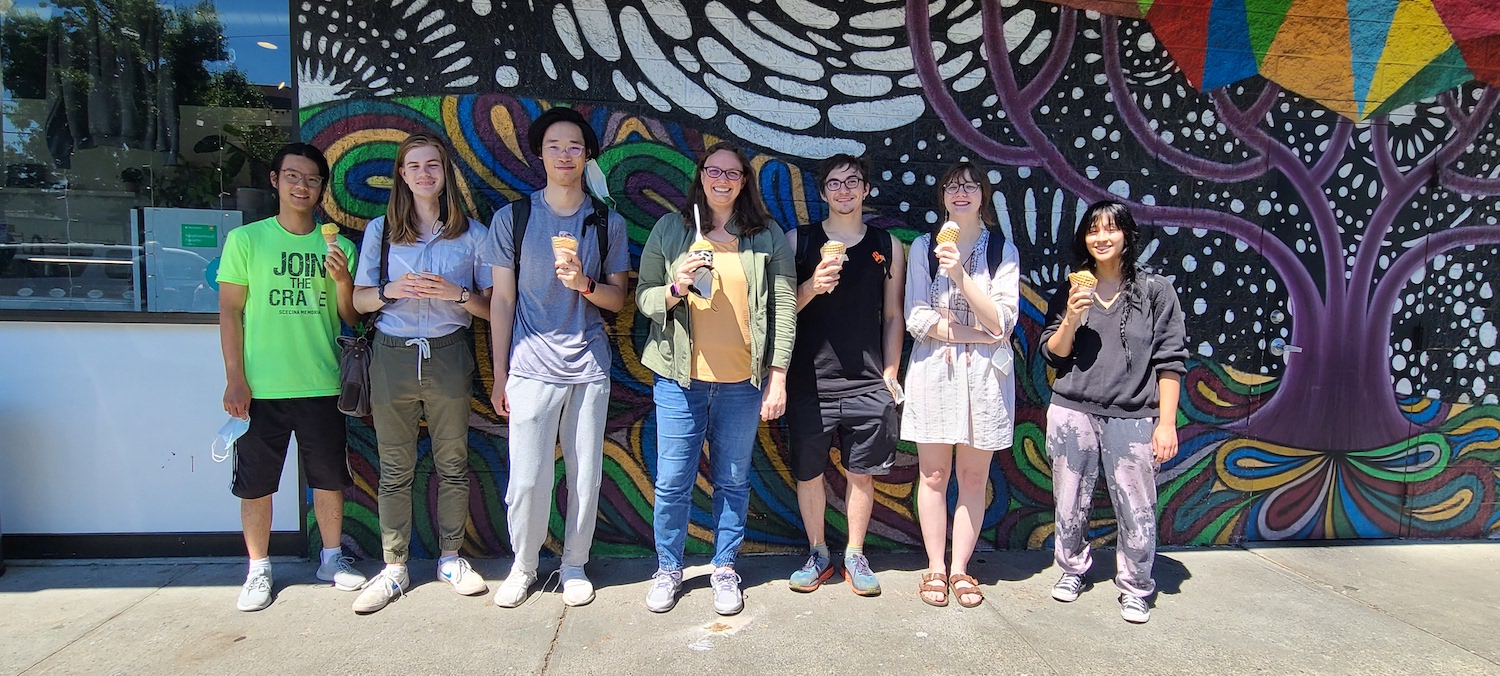
2022 group taking advantage of hot weather + ice cream. From left to right: Li Yi, Henry Jacques, Larry Zeng, Anna, Max Bennett, Nina Young, Alex Richter.

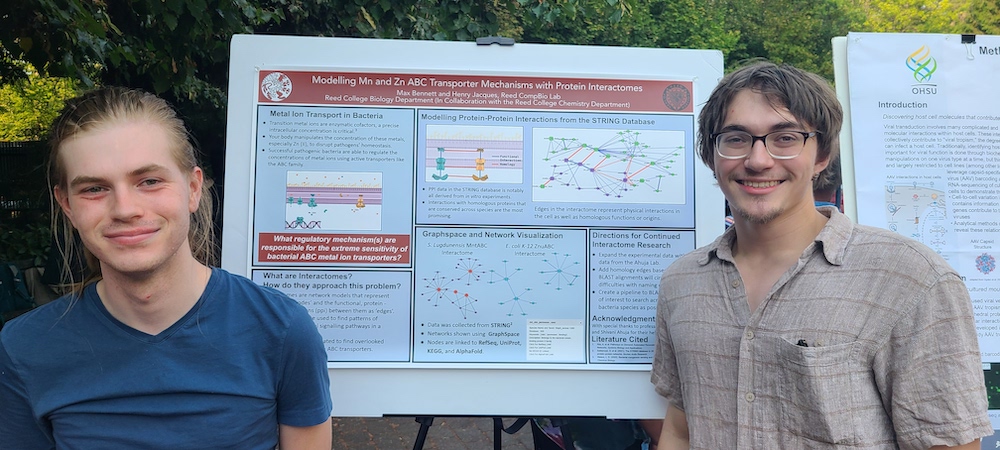
2022 group members presenting at the Reed Fall Poster Session. Left: Li Yi and Hannah Kuder; Right: Henry Jacques and Max Bennett.
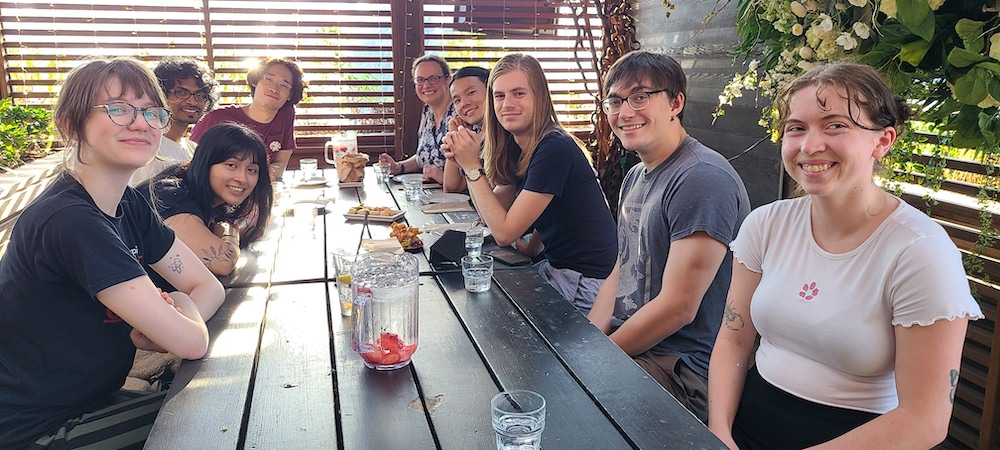

2022 Group Dinner! From left to right: Nina Young, Alex Richter, Pramesh Singh, Larry Zeng, Anna, Li Yi, Henry Jacques, Max Bennett, Hannah Kuder.
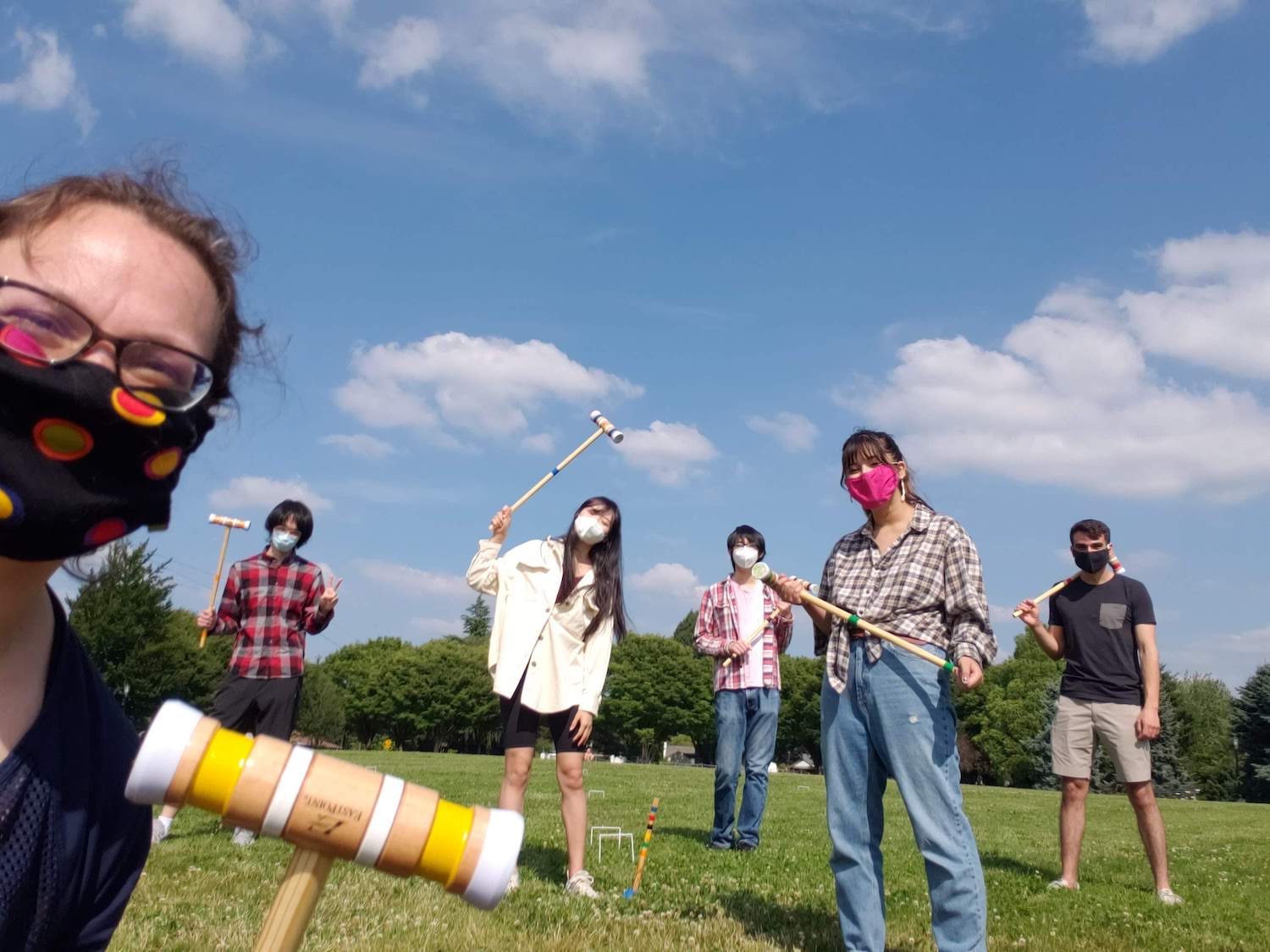

2020 was mostly a mostly remote summer, except for an excellent croquet match. From left to right: Anna, Larry Zeng, Jiarong Li, Frank Zhuang, Alex Richter, Gabe Preising.
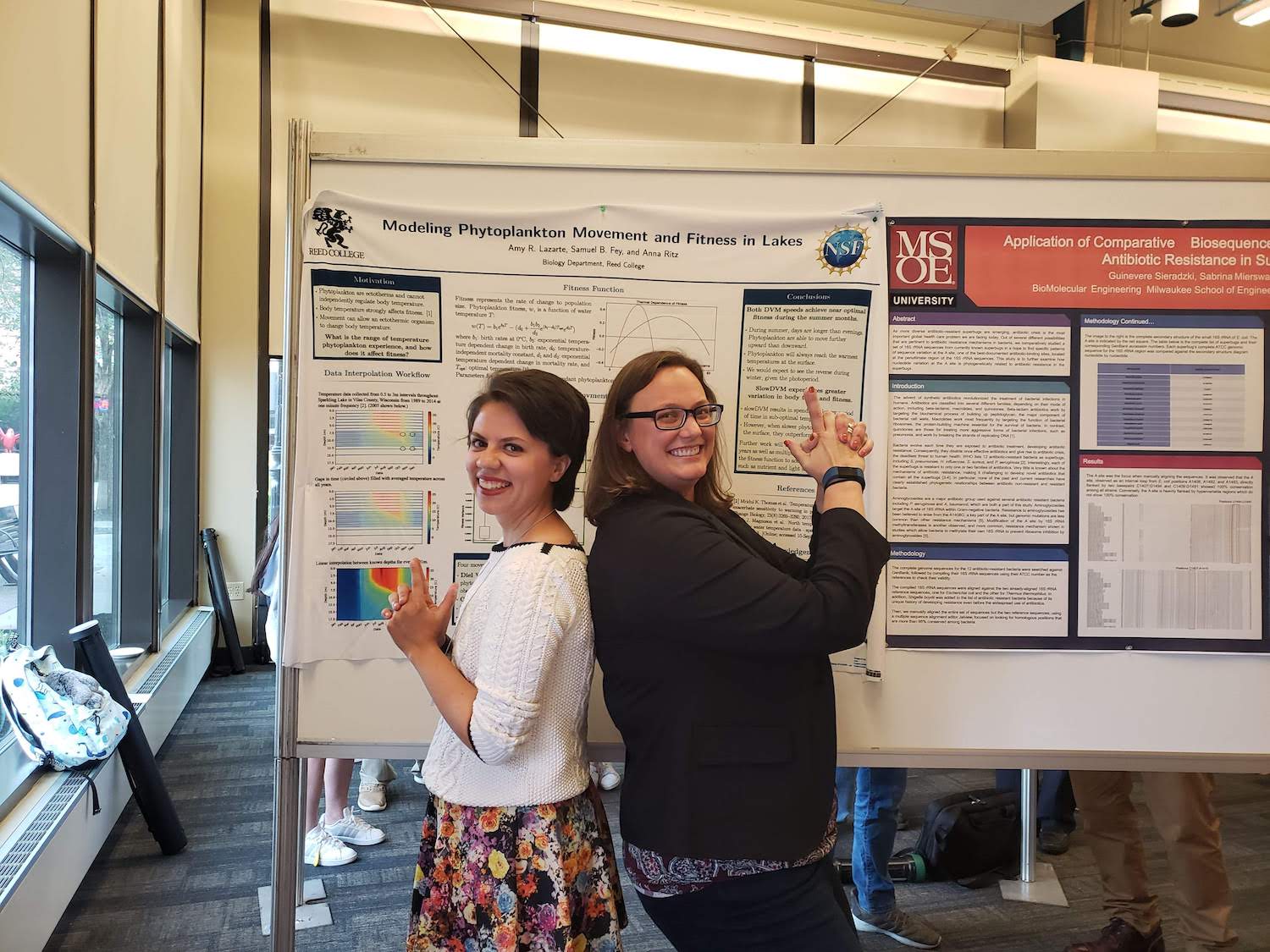

Amy Rose Lazarte presents her thesis work at the 2019 ACM-BCB Conference in Niagara Falls, NY.
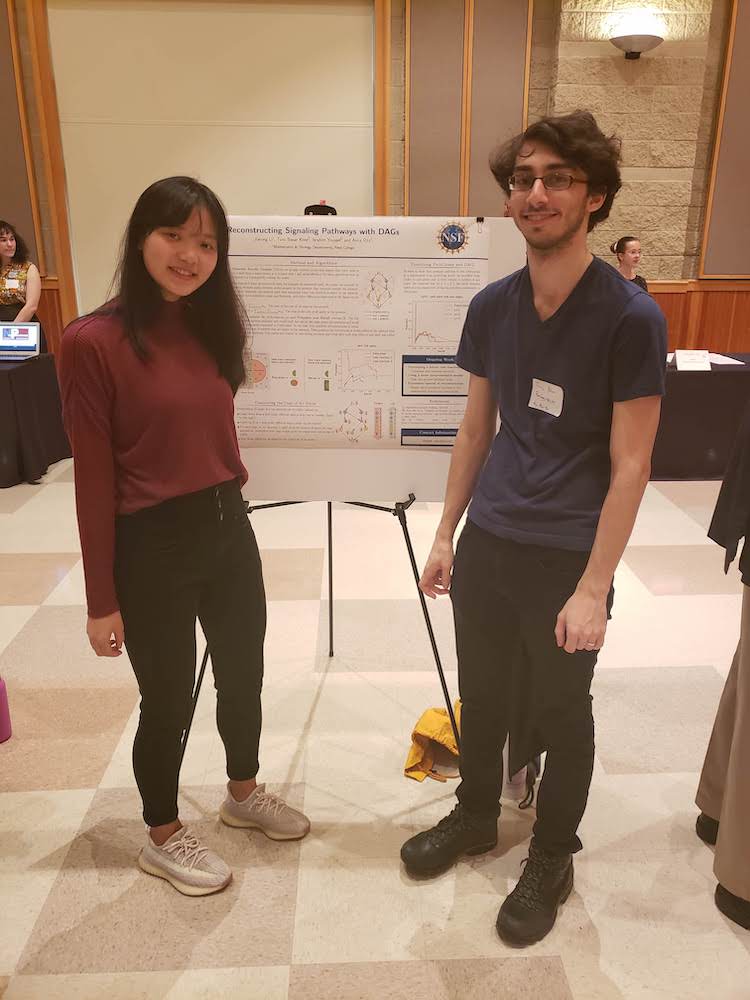
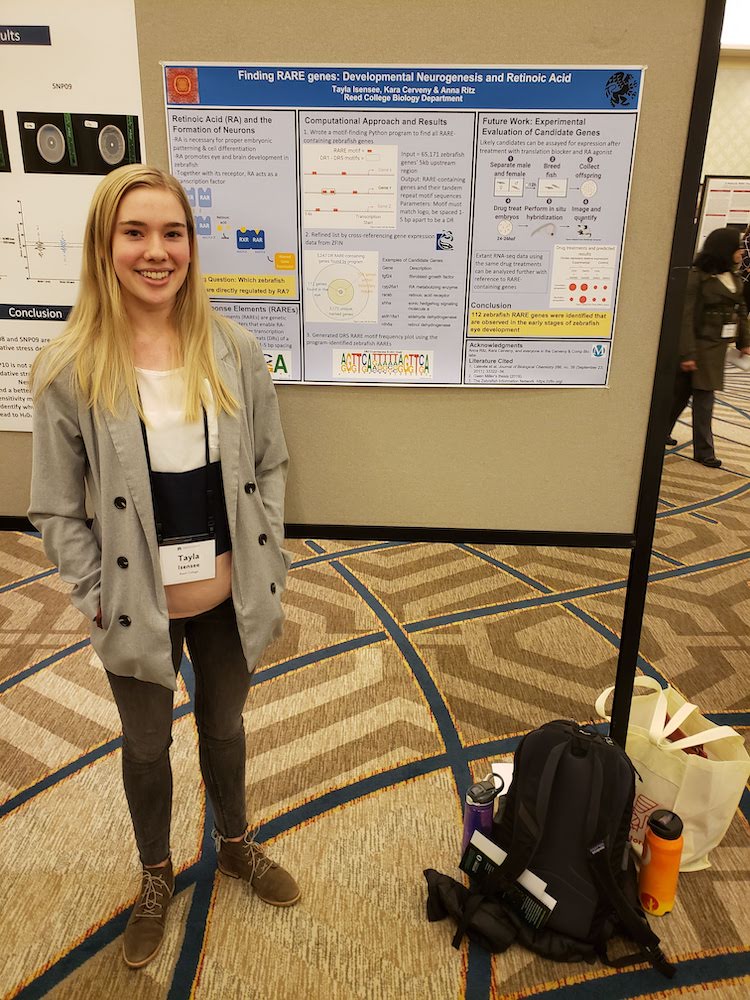

2019 posters presented a local venues. Left: Tayla Isensee presents at the 2019 Murdock Trust Undergraduate Science conference. Right: Jiarong Li and Tunc Kose present at Reed College President Audrey Bilger's welcome event.
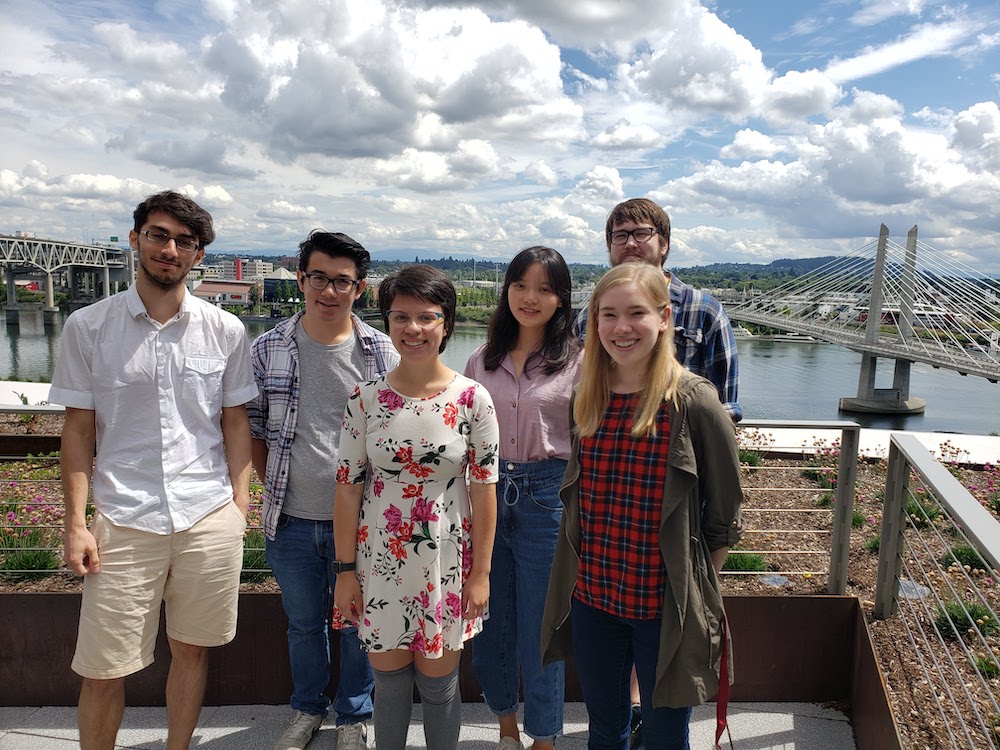

2019 visit to OHSU's Knight Cancer Research Center. From left to right: Tunc Kose, Alex King, Amy Rose Lazarte, Jiarong Li, Tayla Isensee, and Karl Young.

2017 Hike up Hamilton Mountain with Suzy Renn's group and other students (we're tiny!).

2016 Thesis student photo! From left to right: Heather Milne, Nicole Ezell, Barney Potter, and Anna Ritz.
The CompBio Lab
Located in Biology B203, the Computational Biology (CompBio) lab is a resource for Biology students working on computationally-intensive research and thesis projects.

The lab (shown here in October 2015) now includes machines and a printer, in addition to the large display and whiteboard.
Machines
B203 houses Linux computers, including eight quad Intel Core i7-4790 processors with 16GB memory and 256GB solid state hard drives. Additionally, we have one Dual Intel Xeon E5-2680 v4 Processor (14 cores) with 128GB memory and a 512GB solid state hard drive with an extra 2TB serial-ATA hard drive for storage.
Students using the CompBio Lab will become familiar with unix-like environments and working from a terminal. Here are some places to get started.
- Reed's Computer User Services has a Unix help page with many common commands and tips.
- This amazing command-line bootcamp walks people through tutorials, all within a web browser.
- There are many nice command-line tools for manipulating text files.
- grep for matching strings and patterns in text.
- awk for rearranging columns of text.
- sed for searching and replacing strings and patterns in text.
- Use top to see the processes currently running on your machine.
- To see the size of a file (or files in a directory), use ls:
ls -lh - To see the size of the contents of an entire directory (e.g., mydir), use du:
du -h --max-depth 1 mydir
Access
B203 requires key-card access. By default, all of my thesis students will get access. If you are working on a computationally-intensive project, talk to me to see if it's appropriate for CompBio Lab use.
Use your username and password to log onto the machines. Just like the computer labs in the ETC, any files saved on the local machine are erased when you log out. Use file storage systems such as AFS or Google Drive to save your work (check out Sharing at Reed for a list of options. Save early, save often. Talk to me if you begin working with very large files that make this prohibitive.
You may choose to manage your projects with a versioning system such as GIT or SVN. GitHub is a popular place for hosting and managing projects (though they have a maximum upload size limit so storing data on GitHub is not ideal).
- Git - the simple guide is a one-page quick setup tutorial.
- The GitHub Cheat-Sheet is available in other languages.
- GitHub Guides offers tutorials on common workflows.
- TryGit is an interactive, example-based tutorial.
- These and dozens of other tutorials are listed on GitHub Help.
Remote Access
The machines can be remotely accessed when students are off campus. Only students with swipe access can remotely log into the machines. The machines are labeled bioinf1 to bioinf8.
You must first fill out a Virtual Private Network (VPN) request form, available on the VPN Policy Page. If you want to use your personal laptop for remote access, you may need to take extra security measures in order for Information Technology to grant you access.
To log into a particular machine (e.g., bioinf5), you can ssh with
ssh username@bioinf5.reed.edu
where username is your Reed login. You will be prompted for your password.
Installed Packages
To request other software, email me with the name and download page. After I check the license, I will pass it on to Computer User Services and it will be installed on the machines within a couple days. To request Python modules or R packages, simply give me the module/package name.
Software:- blasr
- Long read aligner designed for data sequenced by Pacific Biosciences (PacBio). [use cases]
- blat
- BLAST-like alignment tool. [user guide]
- ncbi-blast+
- Suite of Basic Local Alignment Search Tools (BLAST). [user manual]
- synapseclient
- Provides an interface to Synapse, a collaborative platform for data-intensive computational biology projects. [Python client]
- inkscape
- Free and open-source vector graphics editor (think Adobe Illustrator without the price tag). [tutorials]
- samtools
- Utilities for the Sequence Alignment/Map (SAM) format. [manpage]
- matplotlib
- A 2D plotting library. [gallery with source code]
- networkx
- A software package for the creation, manipulation, and study of networks. [tutorial]
- adgenet
- Multivariate Analysis of Genetic Markers. [reference manual]
- ape
- Analyses of Phylogenetics and Evolution. [reference manual]
- Rsamtools
- Utilities for Sequence Alignment (R package). [documentation]