Research
PET plastic degradation
With the growing problem of plastic pollution, both on land and in our oceans, we see the biodegradation of these waste products to be an important part of the solution to this global issue. For her senior thesis Morgan Vague (Reed ’18) isolated three Pseudomonas and two Bacillus spp. that degrade polyethylene terephthalate, or PET plastic, the material used for disposable water bottles. We want to gain a greater understand the biodegradation of PET by these bacteria, both biologically and chemically, to make the process more rapid. The genome sequences of the five bacteria, and evidence that they degrade PET plastic in a synergistic manner recently was published in Microbiology Resource Announcements. Morgan Vague's TED Talk describing the isolation of the bacteria has over 2 million views. The audio book, titled Using Hungry Microbes to Devour Plastic Pollution, can be found on the science communication channel SciPod. Outreach on the science communication channel, SciPod. This work is funded by NSF grants #1931150 and #2246498.
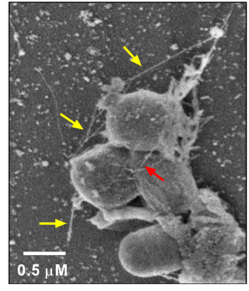
Pseudomonas and Bacillus bacteria forming a biofilm on PET plastic. Pili formation (yellow and red arrows) by bacteria aid in adherence and biofilm formation (Morgan Vague, Senior Thesis, 2018). Image courtesy of Claudia S. López, PhD, Director Multiscale Microscopy Core at OHSU.
Biorecycling
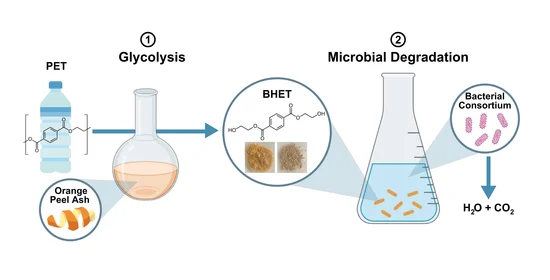
Nearly 600 billion single-use plastic water bottles are produced every year, of which only a small percentage are recycled. This portion of the project focuses on a novel recycling process to decrease the plastic waste that is currently threatening our planet. We are using a two-step, biologically friendly method to degrade PET water bottles, where terephthalic acid can be recovered from culture media and reused to polymerize new PET products. While we predict that complete degradation is achievable using this method, further experimentation with the bacterial consortium can allow for more circular PET recycling, creating a low-cost technology.
Lab Folks 2025

Senior Research Associate: Sabrina Edwards
Thesis students: Cass Biles, Chloe Hsy, Kate Mitchell, Patricio Palomino, Madelyn Tarara, and Lucas Walker