|
|

|
Chapter
4 – Electrostatic Potentials
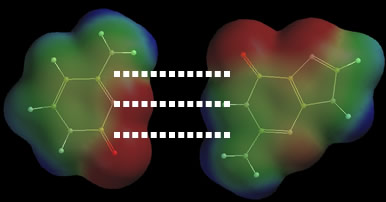
Hydrogen bonds One of the most important nonbonded interactions that can be studied with potential maps is the hydrogen bond or H bond. This kind of “bond” occurs when two atoms, a positively charged hydrogen atom and a negatively charged non-hydrogen atom, interact electrostatically. Hydrogen bonds occur most often between neutral molecules. The interacting atoms in these molecules carry fractional charges, and this type of hydrogen bond can be considered a type of dipole-dipole interaction. The fractionally charged atoms responsible for most hydrogen bonds normally generate significant “local” potentials. Experience shows that hydrogen bonds typically involve a hydrogen atom whose “local” potential is ³ +40 kcal/mol, and a non-hydrogen partner whose “local” potential is £ –40 kcal/mol. Therefore, a survey of the potential maps in this chapter will reveal which atom combinations are most likely to form hydrogen bonds. Hydrogens bonded to either oxygen or fluorine typically generate “local” potentials of +40 kcal/mol or more. These atoms always form strong hydrogen bonds with a suitable partner. Hydrogens bonded to certain types of nitrogen can also generate large “local” potentials, and many hydrogen-bonded systems are based on this type of hydrogen (see below). Hydrogens bonded to carbon rarely generate large potentials, and CHn groups rarely, if ever, form strong hydrogen bonds. On the other side of the ledger, only nitrogen and oxygen atoms seem to generate large negative “local” potentials in neutral compounds. Therefore, most hydrogen bonds look like O–H•••X or N–H•••X, where X = N or O. (Note: “–“ and “•••” correspond to covalent and hydrogen bonds, respectively. The covalent bond is always short and strong, and the hydrogen bond is always long and weak.) An example of the O–H•••O pattern can be found in a cup of water. The two oxygen atoms belong to different water molecules, and we can draw the hydrogen bonded water molecules as HO–H•••OH2 (Ege, p. 31-32). Several examples of the N–H•••X pattern appear in double-stranded DNA. Each DNA strand contains four types of rings (called nucleotides) that are responsible for forming hydrogen bonds with rings in the other strand. The hydrogen bonding capacity of each nucleotide also controls DNA synthesis, and insures that a newly synthesized strand contains the proper sequence of nucleotides. Two nucleotides, cytosine and guanine, can form three hydrogen bonds simultaneously. The other nucleotides, adenine and thymine, can form two (Ege, p. 901). All five hydrogen bonds follow the N–H•••X pattern, and potential maps show that the potentials near these hydrogens are all greater than +40 kcal/mol. Potential maps of cytosine and guanine are shown below. The molecules have been pushed apart so that their respective potential maps can be compared. The maps show that all three of the N–H groups responsible for hydrogen bonds generate large “local” positive potentials. The maps also show that the nitrogen and oxygen partners generate large “local” negative potentials. Each molecule generates a complementary pattern of positive and negative potentials so that, when they approach, three stabilizing interactions will be created.
|