Comparative Functional Genomics
Biology 431 Spring 2008
April 10th 2008
Christine Deyo and Melissa Zarr
Review:
Tirosh, I., Bilu,
Y., and Barkai, N., (2007) Comparative biology: beyond
sequence analysis. Current Opinion in Biotechnology 18:371-377.
Primary Literature:
McCarroll, S.A.,
Murphy,C.T., Zu, S. Pletcher, S.D., Chin, C.S., Jan, Y.N., Kenyon,
C. Bargmann, C.I., and Li, H. (2004) Comparing Genomic expressionpatterns
across species identifies sharerd transcriptional profile in aging. Nature
Genetics 36:197-204.
The topic today is truly Comparative Functional Genomics. The primary paper compares genomic expression patterns across flies, worms, yeast and humans as each organism ages. In order to do this, it was first necessary to identify orthologs across these different species, and identify experimental paradigms that were in some way comperable. The review paper deals with available methods for the identification of "orthologs" or how to define genes that share common function and/or common sequence.
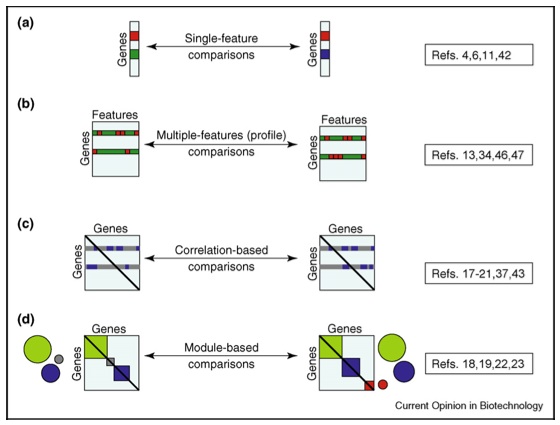
Types of comparative analyses: from single-features to whole networks comparisons. Comparative analysis can be performed with different approaches depending on the type of data and choice of methods. Four different approaches are illustrated with increasing complexity; in each case, squares represent the datasets for two organisms, colors represent different values, and examples are given for conservation (top) and divergence (bottom).
The primary paper used their own, as well as publically available microarray data sets in order to compare gene expression data. Gene Ontology annotations were used to analyze modules of gene expression for functional significance.
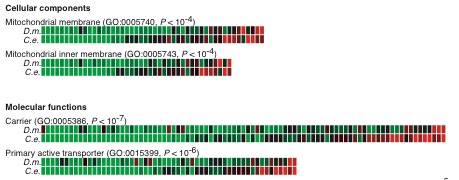 Each
box represents a single gene for flies (top row) and worms
(bottom row). All Genes annotated to a given GO term are shown. Decreased
expression correlated with age is shown in green and increased
is red.
Each
box represents a single gene for flies (top row) and worms
(bottom row). All Genes annotated to a given GO term are shown. Decreased
expression correlated with age is shown in green and increased
is red.
For these 4 terms a decrease in expression was observed
statistically more often than would be expected by chance. Notice that
orthologs (pairs in top and bottom row) tend to be regulated in a similar
fashion in the two very different organisms.
drinks and treats by Will McNitt
who-is-who by Suzy Renn
see this link to an
odd opinion piece by one of the authors.